Can You Identify Mother Baby Pairs Through Oral Bacteria
Abstract
The onset and progress of dental caries and periodontal disease is associated with the oral microbiome. Therefore, it is of import to empathize the factors that influence oral microbiome formation. One of the factors that influence oral microbiome formation is the transmission of oral bacteria from parents. However, it remains unclear when the manual begins, and the difference in contributions of father and mother. Here, we focused on the oral microbiome of 18-calendar month-old infants, at which age deciduous dentition is formed and the oral microbiome is likely to get stable, with that of their parents. Nosotros nerveless saliva from forty 18-month-quondam infants and their parents and compared the diversity and composition of the microbiome using adjacent-generation sequencing of 16S rRNA genes. The results showed that microbial diversity in infants was significantly lower than that in parents and composition of microbiome were significantly dissimilar between infants and parents. Meanwhile, the microbiome of the infants was more similar to that of their mothers than unrelated adults. The bacteria highly shared betwixt infants and parents included not only commensal leaner but likewise disease related bacteria. These results suggested that the oral microbiome of the parents influences that of their children aged < 18 months.
Introduction
Dental caries and periodontal disease are the most prevalent diseases in the worldone,2. These are infectious diseases acquired by oral bacteria and are associated with a shift from symbiotic microbiota to dysbiosis3,iv. Later nascency, normal oral microbiome is formed, which has a symbiotic human relationship with the hostiv. Therefore, information technology is important to sympathise the formation process of normal oral microbiome and to maintain its state for the prevention of disease. However, the process of normal oral microbiome formation after birth and the factors that influence its formation remain unclear.
At nascency, few bacteria are present in the oral cavity5,half dozen. Diversity of the oral microbiome increases over time7. Formation of the oral microbiome in infancy is influenced by breastfeeding8,ix, tooth eruption10, and introduction of solid foodsxi. Moreover, at the age of 18 months, the oral microbiome is known to become stable and the microbial diversity of tongue is comparable to that of adultsvii,12. Also, vertical transmission of mutans streptococci from parents occurs betwixt the ages of xix and 31 monthsthirteen,xiv,15. This menstruation is called the "window of infection"16. Therefore, the transmission of oral bacteria from parents to kid is an important factor influencing the formation of oral microbiome in childhood, and information technology is considered to exist ane of the factors that changes the oral microbiome into a dysbiotic condition related to early childhood caries onset. However, studies of oral bacteria during this period are often limited to cariogenic bacteria such as Streptococcus mutans, and the similarity of the overall microbiome, including commensal bacteria, between parents and child is unknown. Moreover, many reports focus just on mothers and children, and few reports address fathers and children, leaving this latter relationship unclear. A study that involves unabridged families—mothers, fathers and children—will provide a more complete agreement of how the oral microbiome forms before the "window of infection".
In this report, nosotros focused on the oral microbiome of eighteen-calendar month-one-time infants, at which age deciduous dentition is formed, and the oral microbiome becomes stable, with that of their fathers and mothers. We nerveless saliva from xl eighteen-calendar month-onetime infants and their parents and compared the multifariousness and composition of the microbiome using next-generation sequencing of 16S rRNA genes.
Results
Comparison of oral microbial diversity betwixt infants and adults
We nerveless saliva from twoscore groups of xviii-calendar month-old infants and their fathers and mothers. Saliva samples were candy for DNA sequencing and the 16S rRNA region was sequenced using next generation sequencer. Starting time, we compared the diversity of their microbiome. Numbers of detected operational taxonomic units (OTU) and the Shannon diverseness alphabetize of infant grouping were significantly lower than those of the parents grouping (father and mother). No significant differences were observed between the father group and the female parent group (Fig. 1, Steel-Dwass test).
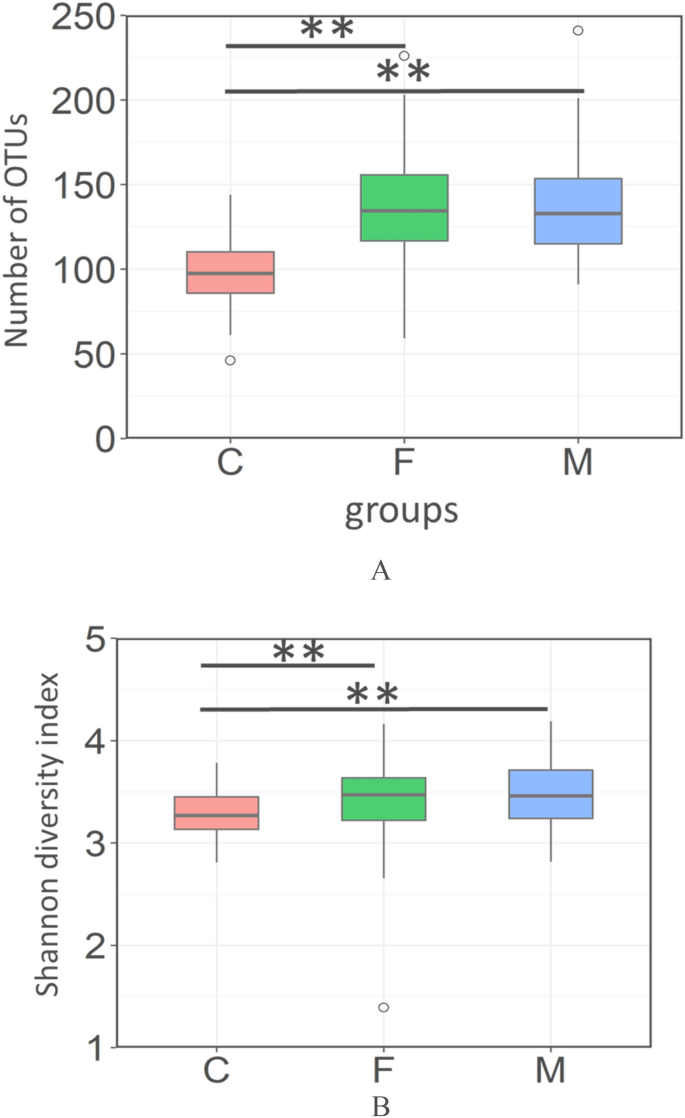
Alpha-diversity of the oral microbiome in each group. Boxplots show the number of observed OTUs (A) and the Shannon diversity index (B) of each group with the number of sequences rarefied to 3000 reads per sample. C; children, F; fathers, M; mothers. Statistically pregnant differences are marked with asterisks (Steel–Dwass test, *p < 0.05, **p < 0.01).
Comparison of oral microbiomes between infants and adults
Subsequently, bacterial composition at the phylum level between xviii-month-old infants, father and mother groups were compared. Amidst the predominant phylum with a > 1% hateful relative abundance in each groups, the relative abundance of Proteobacteria and Fusobacteria were significantly higher in infants, whereas the relative affluence of Bacteroidetes was significantly higher in parents (Fig. 2, Steel–Dwass examination). As with the microbial diversity, meaning differences between father group and mother group were not observed at the phylum level.
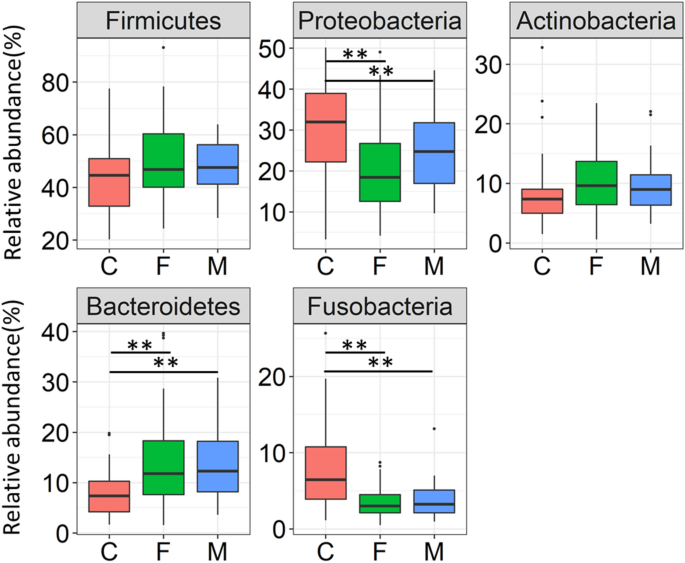
Relative affluence of 5 predominant phyla in the 3 groups. The ascendant phylum showing more than 1% of the hateful relative abundance in the children (C), father (F), and mother (M) groups are shown. Statistically meaning differences are marked with asterisks (Steel–Dwass exam, *p < 0.05, **p < 0.01).
Similarity of microbiomes between infants and adults
The principal coordinate analysis (PCoA) plot based on the UniFrac Altitude metric was performed to compare similarities of microbiome between infants and parents (Fig. iii). A permutational multivariate assay of variance (PERMANOVA) confirmed a meaning deviation between infants and adults when using either Weighted or Unweighted Distance (p < 0.001).
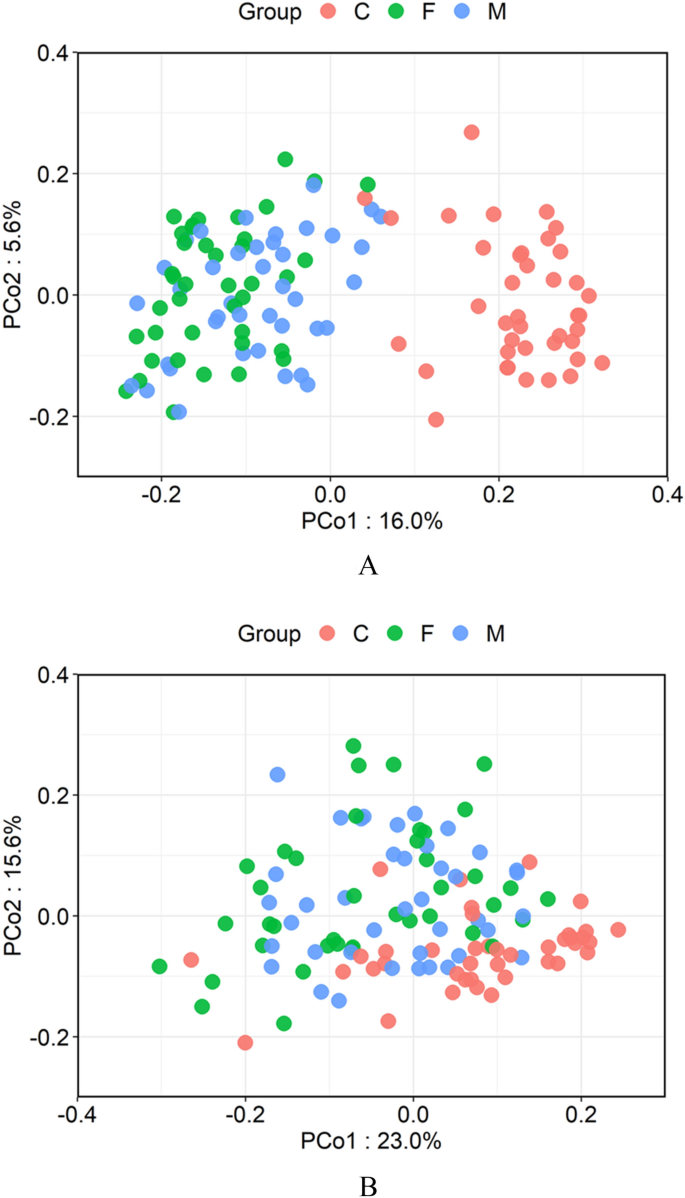
Comparing of the salivary microbiota of the children, fathers, and mothers. (A) Unweighted UniFrac-Principal coordinate analysis (PCoA) and (B) weighted UniFrac-PCoA of salivary microbiota from the three groups. Samples from children, fathers, and mothers are shown as red, green, and blueish, respectively.
Subsequently, to investigate whether an infant's microbiome is more like to their parents than unrelated adults, comparisons of UniFrac Distance betwixt infants and their parents (begetter and mother) or the parents of another infant were performed. The similarity between infants and their mothers was significantly higher than the similarity of infants and unrelated female adults in Unweighted and Weighted UniFrac Distance (Fig. 4, p-values for both Weighted and Unweighted were 0.03). The similarity between infants and their fathers was also college than that of infants and unrelated male adults, simply differences were non significant (Weighted: p = 0.15, Unweighted: p = 0.14).
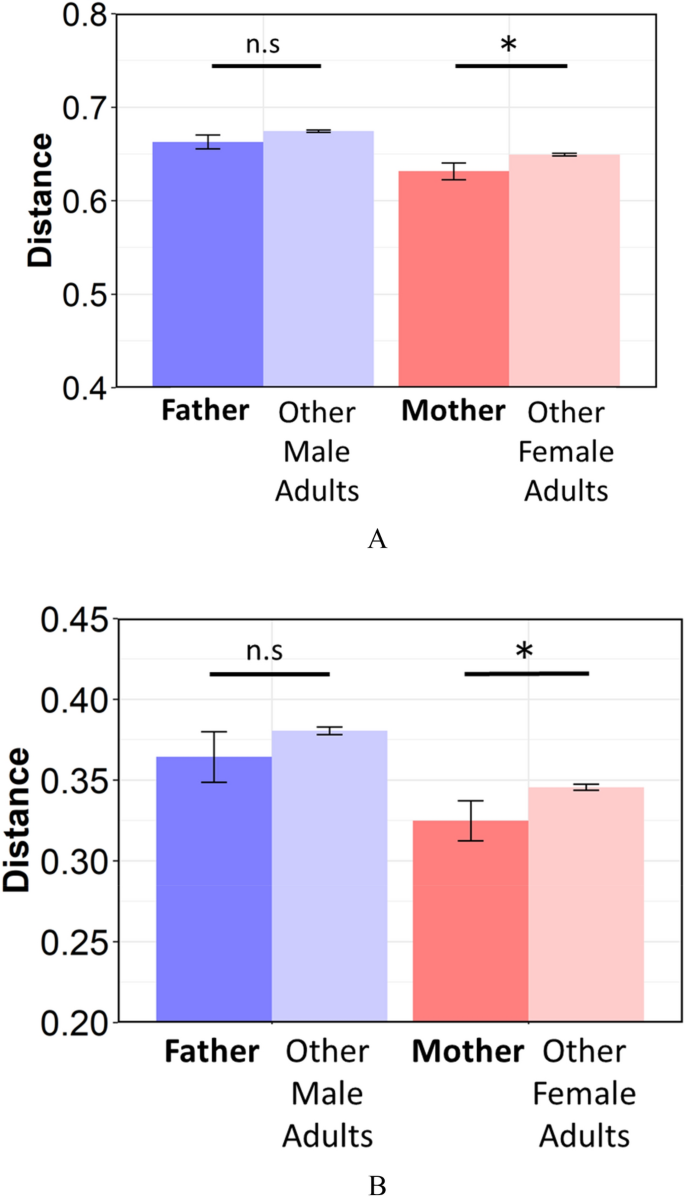
The metrics of (A) unweighted, and (B) weighted Unifrac Altitude between infants and their parents or unrelated adults. Father; between infants and their fathers, other male adults; between infants and unrelated male person adults, Female parent; betwixt infants and their mothers, other female adults; between infants and unrelated female adults. Significant differences are marked with asterisks (U-test, *, p < 0.05 **, p < 0.01) n.s. not significant.
In addition, to examine the relationship of oral microbiome betwixt spouses, we compared the UniFrac Distance betwixt spouses and between unrelated adults. When using either Weighted or Unweighted UniFrac Distance, oral microbiota between spouses showed significantly higher similarity compared with same- or opposite-sex unrelated adults (Supplementaly Fig. S1, p-values for both Weighted and Unweighted were less than 0.01). Farther, no significant differences in the distance between adults of the aforementioned sex or of the reverse sex were found. Thus, no gender differences in adult oral microbiomes were identified.
OTUs shared by infants and their parents
Finally, to make up one's mind whether the bacteria detected in the infants were likewise more oft nowadays in their parents than in unrelated adults, the ratio and total abundance of OTUs shared by the infants and their parents or unrelated adults were compared (Fig. 5). Similar to the UniFrac altitude results, the ratio and total abundance of OTUs shared by infants and their mothers were significantly higher than those shared by infants and unrelated female adults (rate of sharing OTUs: p = 0.003, total abundance of sharing OTUs: p = 0.04). On the other hand, the ratio and full affluence of OTUs shared by infants and their fathers were non significantly higher than those shared by infants and unrelated male adults (rate of sharing OTUs: p = 0.16, full abundance of sharing OTUs: p = 0.11). OTUs highly shared between baby and their parents, were assigned to genus such as Granulicatella, Streptococcus, Veillonella, Neisseria, Haemophilus, Rothia, and Fusobacterium. Table 1 shows a list of OTUs that are highly (> 80%) shared between infants and their parents. Furthermore, OTUs assigned to typical cariogenic and periodontal pathogens (S. mutans and Scarlet Circuitous (Porphyromonas gingivalis, Tannerella forsythia, and Treponema denticola)) were not detected in infants, with the exception of S. mutans which was detected in only ane infant (Supplementary Table S1).
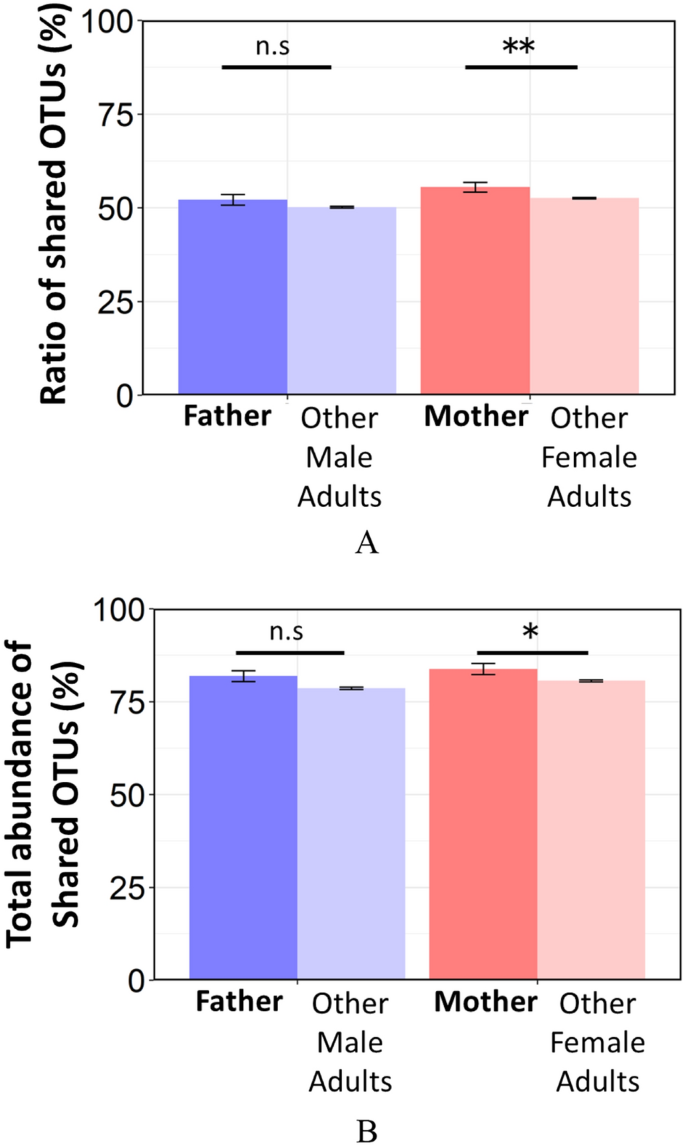
(A) Ratio and (B) full abundance of OTUs detected in the infant that are shared with their parents or unrelated adults. Male parent; between infants and their fathers, other male person adults; betwixt infants and unrelated male person adults, mother; between infants and their mothers, other male person adults; betwixt infants and unrelated female adults. Significant differences are marked with asterisks (U-test, *, p < 0.05 **, p < 0.01). n.s. not significant.
Subsequently, the relationship between indicators known to be related to oral microbiome germination, such as the introduction of infant food, the consecration of deciduous molar eruption, and the mode of feeding (chest milk/artificial milk), and the similarity of the oral microbiome between the infants and parents were analyzed, and none of the indexes exhibited a clear relationship (Supplementary Tabular array S2). Moreover, whether or not the baby was weaning did not affect the similarity between the infant and their parents (Supplementary Table S3). Circadian rhythm has too been described to contribute to the salivary microbiome17; all the same, we did not see significant human relationship betwixt the sampling fourth dimension and similarity of the microbiome between the infants and their parents (Supplementary Table S2).
Give-and-take
In this study, we attempted to clarify the relationships of the oral microbiome in 18-month-onetime infants and their parents. Nosotros collected saliva from forty xviii-month-old infants and their parents and compared diversity and composition of their microbiome using adjacent-generation sequencing of 16S rRNA genes. We showed that diversity and composition of oral microbiome in 18-month-old infants differs from their parents. Meanwhile, the oral microbiome of the infants was more similar to their mothers than that of unrelated female adults. The bacteria highly shared between infants and parents included not only commensal bacteria but also disease related bacteria. These results suggested that the oral microbiome of the female parent influences the oral microbiome of their children by 18 months.
In this study, the oral microbiota of 18-month-onetime infants were less various than that of adults (Fig. 1). This result was consistent with studies comparing the salivary microbiome in infants and adults, from the infants' birth to five years of agexviii,19. These results suggested that the oral microbiome of the infants was still immature at xviii months of age. Moreover, another written report reported that the microbial diversity of the natural language in xviii-calendar month-old infants is comparable to that of adults7. These results suggest that the maturity of the oral microbiome varies depending on the location in the oral cavity.
Significant similarity in microbiomes was found in infants and their mothers compared with unrelated adults (Fig. 4). The ratio and full affluence of OTUs shared by infants and their mothers were significantly higher than those shared by infants and unrelated female adults (Fig. 5). Considering these results, the oral microbiome of parents influences the oral microbiome of infants and both commensal and pathogenic oral bacteria may be transmitted from parents to their infants before the "window of infection".
Information technology has been reported that oral crenel microbiota differs between caries active and healthy children, and the shift from symbiotic microbiome to dysbiotic microbiome is linked to early childhood caries onseteighteen,twenty,21. Regarding the factors causing the shift to dysbiotic microbiome, an increased frequency of saccharide intake is mainly focused on. However, information technology was besides reported that feeding habits are not significantly different between children with or without severe early babyhood caries21, and the transmission of oral bacteria from parents is likewise considered to be i of the of import factors for oral microbiome shift22,23. In this study, most of the leaner highly shared by infants and their parents were oral commensal bacteria such as Streptococcus, Veillonella, Neisseria, Haemophilus, and Rothia. However, OTUs assigned to Fusobacterium nucleatum, which is involved in periodontal disease24,25, were too shared at a high rate. (Table ane). F. nucleatum has been suggested to play an important office in multispecies dental biofilm formation due to its power to attach to a very large variety of different microorganisms26. Consequently, F. nucleatum is considered to exist associated with the transition from a commensal community to a pathogenic community27,28. Thus, the oral microbiome of parents may influence the shift of oral microbiome of their children from symbiotic to dysbiotic microbiome.
In dissimilarity, no significant similarity was detected between infants and their fathers compared with the baby and unrelated adults, suggesting that mothers' oral microbiota take a greater issue on the formation of their children's oral microbiome than fathers'. This finding may exist due to more intimate contact with the mother's microbiome, such as during breastfeeding.
In addition, oral microbiome between spouses was significantly more than like than microbiomes between unrelated adults (Supplementaly Fig. S1). This result suggests that fifty-fifty betwixt individuals that are genetically dissimilar, like oral microbiomes are formed likely by activities such as kissing and contact with aforementioned microbial sources or diets. Kissing causes a temporary exchange of oral bacteria29 and Porphyromonas gingivalis, a typical periodontal pathogen, is known to be transmitted between spousesxxx,31,32. Although we did non obtain lifestyle information such as kissing from the subjects in this study, it is only assumed that spouses' physical contact is one explanation for similarity of oral microbiome betwixt spouses.
The limitation of this report is that analysis using 16S rRNA is not suitable for detailed bacterial type identification. More detailed analysis, such equally shotgun metagenomics, must be used to decide if transmission of oral bacteria occurs among family members. In this report, we could not obtain data on the history of caries, periodontal disease, smoking, or systemic diseases (such as inflammatory bowel disease) that are known to impact the oral microbiome33,34. Therefore, further studies are warranted to understand the effects of these factors on the similarity of the oral microbiome between infants and their parents.
In determination, a comparison of oral microbiomes of 18-calendar month-erstwhile infants and their parents revealed that oral microbiomes were significantly similar betwixt infants and their mothers compared with unrelated adults. The bacteria highly shared between infants and parents included not only commensal bacteria but also disease related leaner. These results suggested that the oral microbiome of the parents influences that of their children earlier the window of infection. Hence, information technology is important for parents to control their oral microbiome past continuous professional treatment and self-intendance, and if they are suffering from oral diseases, should be conscientious of infections in their children.
Methods
Study subjects
We recruited forty eighteen-month-former infants (19 boys and 21 girls) and their biological fathers [anile 26–48 years (mean ± s.d., 34.eight ± iv.9 years)] and mothers [aged 27–42 years (mean ± south.d., 33.2 ± 3.vii years)] who were living together. The fathers and/or mothers were employees or employee's family of a manufacturer located in Tokyo, Japan. Children had non received antibiotics in the six months before the collection of samples. We examined the historic period of introduction of baby food, consecration of deciduous tooth eruption, weaning, and ratio of chest milk/bogus milk. These information are listed in Supplementary Tabular array S4. All adult participants understood the purpose of the written report and provided informed consent. Because our affiliated institutions did not have whatsoever ethics committee, this study was given ethical approval by ethics committee of the academic society (Ethics committee of the Japanese Society for Oral Health, Tokyo, Nihon, Issuing number: No. 26-v). Although authors are members of this bookish society, none of the states are affiliated to its ideals committee. All experiments were performed in accordance with approved guidelines.
Sample drove
Before sample collection, participants were instructed non to brush their teeth from the last meal to the fourth dimension of sampling and were prohibited from eating or drinking for at least 1 h before sampling. Infant saliva was collected using SalivaBio Babe's Swabs (Salimetrics, United states of america, CA)35. Saliva of infants were collected by sucking saliva accumulated in the oral cavity using a swab stick. Parent'southward saliva samples were collected equally mouth-rinsed water36. Briefly, participants rinsed their mouth vigorously with three mL sterilised water for 10 s, and so spat into a sterilised specimen tube. All samples were stored at refrigerated condition and centrifuged at 16,400 ×g for 5 min within 30 h subsequently drove. Resulting pellets were stored at − lxxx °C until DNA extraction.
Deoxyribonucleic acid extraction and sequencing of 16S rRNA gene amplicons
Genomic DNA was isolated from the collected samples using a Nexttec 1-Step Deoxyribonucleic acid Isolation Kit (nexttec Biotechnologie GmbH, Leverkusen, Federal republic of germany). PCR used universal primers (27Fmod and 338R) for 16S rRNA gene sequencing, equally previously described36,37. PCR used Ex Taq polymerase (Takara Bio, Shiga, Japan) and approximately 20 ng of template Deoxyribonucleic acid.
Thermal cycling was performed in a Veriti Thermal Cycler (Life Technologies Japan, Tokyo, Nihon). Cycling atmospheric condition were: initial denaturation at 96 °C for 2 min, followed past 25 cycles of denaturation at 96 °C for 30 due south, annealing at 55 °C for 45 south, extension at 72 °C for 1 min, and final extension at 72 °C. PCR amplicons were purified using AMPure XP magnetic purification chaplet (Beckman Coulter, CA, USA) and quantified using a Quant-iT PicoGreen dsDNA Assay Kit (Life Technologies Japan). After quantification, mixed samples were prepared past pooling approximately equal amounts of each amplified Deoxyribonucleic acid. Samples were sequenced using a MiSeq Reagent Kit V3 (300 × ii cycles) and a MiSeq sequencer (Illumina, CA, Us), following the manufacturer's instructions.
Data processing
Nosotros used an analysis pipeline for processing the 16S rRNA gene V1–V2 region, as previously reported34,38. Briefly, after multiplexed sequencing of the 16S amplicons, sequences were assigned to samples based on their barcode sequences. Reads with an average quality value < 25, inexact matches to both universal primers, and possible chimeric reads were eliminated. Among high-quality reads, 3000 reads per sample were randomly chosen and used for the comparative microbiome analysis. We sorted selected reads with the average quality value and grouped them into OTUs using the UCLUST (five.5.2.32) algorithm with a 97% identity threshold39. Taxonomic assignments for each OTU were made by similarity searching against publicly bachelor 16S database using the GLSEARCH program (v.36.3.8 g). The 16S database was constructed from three publically available databases, as previously described19: Ribosomal Database Projection (RDP) v.10.31, CORE (http://microbiome.osu.edu/ (31 January 2017, appointment last accessed)), and the reference genome sequence database obtained from the NCBI FTP site (ftp://ftp.ncbi.nih.gov/genbank/ (December 2011, date last accessed)). For assignment at the phylum levels, sequence similarity thresholds of lxx% were applied, respectively39. All high-quality 16S V1–V2 sequences were submitted to the DDBJ/GenBank/EMBL database (Accession number DRA010385).
Data analysis
We used Isle of mann-Whitney U-test and Steel-Dwass test where appropriate for comparisons of chiselled variables. We likewise used UniFrac distance40 for dissimilarity (distance) assessment between pairs of samples. PCoA was used to visualise similarities/dissimilarities in microbiome structures from the UniFrac Distance. We conducted a PERMANOVA to compare overall microbiome structures. Differences at p < 0.05 were considered statistically significant. All analyses were performed using R software programme (v3.4.iii).
References
-
Chapple, I. L. et al. Interaction of lifestyle, behavior or systemic diseases with dental caries and periodontal diseases: Consensus study of group2 of the joint EFP/ORCA workshop on the boundaries between caries and periodontal diseases. J. Clin. Periodontol. 44(Suppl 18), S39–S51 (2017).
-
Marcenes, W. et al. Global burden of oral conditions in 1990–2010: A systematic analysis. J. Paring. Res. 92, 592–597 (2013).
-
Marsh, P. D. In sickness and in health-what does the oral microbiome mean to u.s.? An ecological perspective. Adv. Dent. Res. 29, 60–65 (2018).
-
Kilian, M. et al. The oral microbiome—An update for oral healthcare professionals. Br. Dent. J. 221, 657–666 (2016).
-
Rotimi, V. & Duerden, B. The development of the bacterial flora in normal neonates. J. Med. Microbiol. xiv, 51–62 (1981).
-
Nelson-Filho, P. et al. Dynamics of microbial colonization of the rima oris in newborns. Braz. Dent. J. 24, 415–419 (2013).
-
Kageyama, S. et al. Transition of bacterial variety and composition in natural language microbiota during the offset ii years of life. mSphere four, e00187-nineteen (2019).
-
Timby, N. et al. Oral microbiota in infants fed a formula supplemented with bovine milk fat globule membranes—A randomized controlled trial. PLoS I 12, e0169831 (2017).
-
Al-Shehri, S. Southward. et al. Deep sequencing of the 16S ribosomal RNA of the neonatal oral microbiome: A comparison of breast-fed and formula-fed infants. Sci. Rep. 6, 38309 (2016).
-
Mason, K. R., Chambers, S., Dabdoub, Southward. M., Thikkurissy, S. & Kumar, P. Due south. Characterizing oral microbial communities across dentition states and colonization niches. Microbiome six, 67 (2018).
-
Sulyanto, R. One thousand., Thompson, Z. A., Beall, C. J., Leys, East. J. & Griffen, A. L. The predominant oral microbiota is acquired early in an organized pattern. Sci. Rep. 9, 10550 (2019).
-
Li, F., Tao, D., Feng, 10., Wong, M. C. K. & Lu, H. Establishment and development of oral microflora in 12–24 month-old toddlers monitored by high-throughput sequencing. Front end. Cell Infect. Microbiol. 8, 422 (2018).
-
Klein, M. I., Florio, F. Chiliad., Pereira, A. C., Hofling, J. F. & Goncalves, R. B. Longitudinal study of transmission, diversity, and stability of Streptococcus mutans and Streptococcus sobrinus genotypes in Brazilian plant nursery children. J. Clin. Microbiol. 42, 4620–4626 (2004).
-
Li, Y., Ismail, A. I., Ge, Y., Tellez, K. & Sohn, Westward. Similarity of bacterial populations in saliva from African–American mother–child dyads. J. Clin. Microbiol. 45, 3082–3085 (2007).
-
da Silva Bastos, V. A. et al. Mother-to-child transmission of Streptococcus mutans: A systematic review and meta-analysis. J. Dent. 43, 181–191 (2015).
-
Caufield, P. Due west., Cutter, G. R. & Dasanayake, A. P. Initial acquisition of mutans streptococci by infants: Evidence for a discrete window of infectivity. J. Dent. Res. 72, 37–45 (1993).
-
Takayasu, L. et al. Cyclic oscillations of microbial and functional limerick in the human salivary microbiome. DNA Res. 24, 261–270 (2017).
-
Dashper, S. G. et al. Temporal development of the oral microbiome and prediction of early childhood caries. Sci. Rep. 9, 19732 (2019).
-
Lif Holgerson, P. et al. A longitudinal study of the evolution of the saliva microbiome in infants 2 days to 5 years compared to the microbiome in adolescents. Sci. Rep. 10, 9629 (2020).
-
Teng, F. et al. Prediction of early childhood caries via spatial-temporal variations of oral microbiota. Cell Host Microbe 18, 296–306 (2015).
-
Hurley, East. et al. Comparison of the salivary and dentinal microbiome of children with severe-early childhood caries to the salivary microbiome of caries-free children. BMC Oral Health 19, xiii (2019).
-
Childers, N. K. et al. Clan betwixt early childhood caries and colonization with Streptococcus mutans genotypes from mothers. Pediatr. Dent. 39, 130–135 (2017).
-
Law, Five., Seow, Due west. K. & Townsend, G. Factors influencing oral colonization of mutans streptococci in young children. Aust. Dent. J. 52, 93–159 (2007).
-
Moore, W. East. & Moore, 50. 5. The bacteria of periodontal diseases. Periodontology 2000(v), 66–77 (1994).
-
Yang, Due north. Y. et al. Progression of periodontal inflammation in adolescents is associated with increased number of Porphyromonas gingivalis, Prevotella intermedia, Tannerella forsythensis, and Fusobacterium nucleatum. Int. J. Paediatr. Dent. 24, 226–233 (2014).
-
Kolenbrander, P. E. et al. Advice among oral bacteria. Microbiol. Mol. Biol. Rev. 66, 486–550 (2002).
-
Kolenbrander, P. et al. Oral multispecies biofilm development and the fundamental function of jail cell–cell distance. Nat. Rev. Microbiol. viii, 471–480 (2010).
-
Han, Y. Due west. Fusobacterium nucleatum: A commensal-turned pathogen. Curr. Opin. Microbiol. 23, 141–147 (2015).
-
Kort, R. et al. Shaping the oral microbiota through intimate kissing. Microbiome 2, 41 (2014).
-
van Steenbergen, T. J., Petit, One thousand. D., Scholte, 50. H., van der Velden, U. & de Graaff, J. Transmission of Porphyromonas gingivalis between spouses. J. Clin. Periodontol. 20, 340–345 (1993).
-
Asano, H., Ishihara, K., Nakagawa, T., Yamada, S. & Okuda, Yard. Relationship between transmission of Porphyromonas gingivalis and fimA blazon in spouses. J. Periodontol. 74, 1355–1360 (2003).
-
Bennani, M. et al. Shared detection of Porphyromonas gingivalis in cohabiting family unit members: A systematic review and meta-analysis. J. Oral. Microbiol. 12, 1687398 (2019).
-
Wu, J. et al. Cigarette smoking and the oral microbiome in a large study of American adults. ISME J. 10, 2435–2446 (2016).
-
Said, H. S. et al. Dysbiosis of salivary microbiota in inflammatory bowel disease and its association with oral immunological biomarkers. DNA Res. 21, fifteen–25 (2014).
-
Jordan, S. et al. Maternal-child microbiome: Specimen collection, storage and implications for research and practice. Nurs. Res. 66, 175–183 (2017).
-
Jo, R. et al. Comparing of oral microbiome profiles in stimulated and unstimulated saliva, tongue, and mouth-rinsed h2o. Sci. Rep. 9, 16124 (2019).
-
Kim, S. W. et al. Robustness of gut microbiota of salubrious adults in response to probiotic intervention revealed by high-throughput pyrosequencing. DNA Res. xx, 241–253 (2013).
-
Iwasawa, Chiliad. et al. Dysbiosis of the salivary microbiota in pediatric-onset primary sclerosing cholangitis and its potential equally a biomarker. Sci. Rep. 8, 5480 (2018).
-
Urushiyama, D. et al. Microbiome profile of the amniotic fluid as a predictive biomarker of perinatal outcome. Sci. Rep. vii, 12171 (2017).
-
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: An effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2011).
Author information
Affiliations
Contributions
R.J., K.Tsutsumi, Grand.M., Thousand. Takeda, K.South., and S.One thousand. planned the report. R.J., K.Y., Y.A., K. Tsutsumi, C.I., Thou. Takeda, M.Chiliad., and S.M. contributed to the drove of samples. R.J., K.Y., and Y.A. contributed to analysis of the primary sequence data of salivary samples. R.J., and Yard.Y. contributed to the statistical assay of the data. R.J. wrote the showtime draft, and Due east.N., K.Due south. and Southward.M. contributed to the completion of the manuscript. All authors read and approved the concluding manuscript.
Respective author
Ethics declarations
Competing interests
R.J., Thou.Y., Y.A., G. Tsutsumi, C.I., M.M., E.N., and K.S. work at King of beasts Corporation. The other authors declare no competing interests.
Additional information
Publisher's annotation
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Artistic Commons Attribution four.0 International License, which permits apply, sharing, adaptation, distribution and reproduction in whatsoever medium or format, as long as you give appropriate credit to the original author(south) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party cloth in this commodity are included in the article's Artistic Commons licence, unless indicated otherwise in a credit line to the material. If cloth is not included in the article'due south Creative Commons licence and your intended apply is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/four.0/.
Reprints and Permissions
About this commodity
Cite this commodity
Jo, R., Yama, K., Aita, Y. et al. Comparison of oral microbiome profiles in xviii-month-old infants and their parents. Sci Rep 11, 861 (2021). https://doi.org/ten.1038/s41598-020-78295-ane
-
Received:
-
Accepted:
-
Published:
-
DOI : https://doi.org/10.1038/s41598-020-78295-i
Further reading
Comments
By submitting a comment you hold to abide past our Terms and Customs Guidelines. If you find something calumniating or that does not comply with our terms or guidelines delight flag it as inappropriate.
dominguezwitchany82.blogspot.com
Source: https://www.nature.com/articles/s41598-020-78295-1
0 Response to "Can You Identify Mother Baby Pairs Through Oral Bacteria"
Postar um comentário